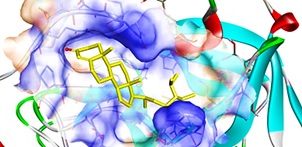
Activity Prediction of Bioactive Compounds Contained in Etlingera elatior Against the SARS-CoV-2 Main Protease: An In Silico Approach
Abstract
The COVID-19 pandemic has become a serious problem today, with its prevalence increasing every day. The SARS-CoV-2 main protease (MPro) is a promising therapeutic target to inhibit replicating and spreading the virus that causes COVID-19. The compounds contained in the Etlingera elatior plant has the potential. This study aimed to examine the compounds' activity in E. elatior against SARS-CoV-2 MPro using in silico methods. A total of seven compounds contained in E. elatior were obtained from the Knapsack database. The compounds were then docked into the SARS-CoV-2 MPro receptor's active site with the PDB ID 6LU7. Afterward, the biological activities were predicted by the PASS prediction webserver. The molecular docking results showed that ergosterol peroxide and sitostenone had the best binding energy with -10.40 kcal/mol and -9.17 kcal/mol, respectively. The in silico PASS prediction showed it has potential as antiviral therapy. It concluded ergosterol peroxide and sitostenone has the potential as SARS-CoV-2 MPro inhibitor candidate.
Full text article
References
Afendi, F.M., Okada, T., Yamazaki, M., Hirai-Morita, A., Nakamura, Y., Nakamura, K., Ikeda, S., Takahashi, H., Altaf-Ul-Amin, M., Darusman, L.K., Saito, K., & Kanaya, S. (2012). KNApSAcK family databases: integrated metabolite-plant species databases for multifaceted plant research. Plant and Cell Physiology, 53(2), e1. doi:10.1093/pcp/pcr165
Bell, E.W. & Zhang, Y. (2019). DockRMSD: an open-source tool for atom mapping and RMSD calculation of symmetric molecules through graph isomorphism. Journal of Cheminformatics, 11(1), 40. doi:10.1186/s13321-019-0362-7
Cherrak, S.A., Merzouk, H., & Mokhtari-Soulimane, N. (2020). Potential bioactive glycosylated flavonoids as SARS-CoV-2 main protease inhibitors: A molecular docking and simulation studies. PLoS One, 15(10), e0240653. doi:10.1371/journal.pone.0240653
Dai, W., Zhang, B., Jiang, X.M., Su, H., Li, J., Zhao, Y., Xie, X., Jin, Z., Peng, J., Liu, F., Li, C., Li, Y., Bai, F., Wang, H., Cheng, X., Cen, X., Hu, S., Yang, X., Wang, J., Liu, X., Xiao, G., Jiang, H., Rao, Z., Zhang, L.K., Xu, Y., Yang, H., & Liu, H. (2020). Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science, 368(6497), 1331-1335. doi:10.1126/science.abb4489
Du, Q.S., Wang, S.Q., Zhu, Y., Wei, D.Q., Guo, H., Sirois, S., & Chou, K.C. (2004). Polyprotein cleavage mechanism of SARS CoV Mpro and chemical modification of the octapeptide. Peptides, 25(11), 1857-1864. doi:10.1016/j.peptides.2004.06.018
Enmozhi, S., Raja, K., Sebastine, I., & Joseph, J. (2020). Andrographolide as a potential inhibitor of SARS-CoV-2 main protease: an in silico approach. Journal of Biomolecular Structure and Dynamics, Online ahead of print, 1-7. doi:10.1080/07391102.2020.1760136
Fristiohady, A., Wahyuni, W., Malik, F., Yusuf, M.I., Salma, W.O., Hamsidi, R., Talebong, F., Yuliansyah, Y., Purnama, L.O.M.J., Saripuddin, S., & Sahidin, S. (2020). Level of Cytokine Interleukin-6 and Interleukin 1-β on Infectious Rat Model Treated with Etlingera elatior (Jack) R.M. Smith Fruit Extract as Immunomodulator. Borneo Journal of Pharmacy, 3(2), 52-57. doi:10.33084/bjop.v3i2.1318
Fristiohady, A., Zubaydah, W.O.S., Wahyuni, W., Mirda, M., Saripuddin, S., Andriani, R., Purnama, L.O.M.J., & Sahidin, S. (2019). Immunomodulator Activity of Effervescent Granule of Wualae Fruit (Etlingera elatior (Jack) R.M. Smith) Based on Specific Phagocytic Activity. Borneo Journal of Pharmacy, 2(2), 35-40. doi:10.33084/bjop.v2i2.868
Ghosh, R., Chakraborty, A., Biswas, A., & Chowdhuri, S. (2020). Potential therapeutic use of corticosteroids as SARS CoV-2 main protease inhibitors: a computational study. Journal of Biomolecular Structure and Dynamics, Online ahead of print, 1-14. doi:10.1080/07391102.2020.1835728
Hui, D.S., Azhar, E.I., Madani, T.A., Ntoumi, F., Kock, R., Dar, O., Ippolito, G., Mchugh, T.D., Memish, Z.A., Drosten, C., Zumla, A., & Petersen, E. (2020). The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health — The latest 2019 novel coronavirus outbreak in Wuhan, China. International Journal of Infectious Diseases, 91, 264-266. doi:10.1016/j.ijid.2020.01.009
Islam, M.T., Sarkar, C., El-Kersh, D.M., Jamaddar, S., Uddin, S.J., Shilpi, J.A., & Mubarak, M.S. (2020). Natural products and their derivatives against coronavirus: A review of the non-clinical and pre-clinical data. Phytotherapy Research, 34(10), 2471-2492. doi:10.1002/ptr.6700
Jin, Z., Du, X., Xu, Y., Deng, Y., Liu, M., Zhao, Y., Zhang, B., Li, X., Zhang, L., Peng, C., Duan, Y., Yu, J., Wang, L., Yang, K., Liu, F., Jiang, R., Yang, X., You, T., Liu, X., Yang, X., Bai, F., Liu, H., Liu, X., Guddat, L.W., Xu, W., Xiao, G., Qin, C., Shi, Z., Jiang, H., Rao, Z., & Yang, H. (2020). Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature, 582, 289-293. doi:10.1038/s41586-020-2223-y
Joshi, T., Joshi, T., Sharma, P., Mathpal, S., Pundur, H., Bhatt, V., & Chandra, S. (2020). In silico screening of natural compounds against COVID-19 by targeting Mpro and ACE2 using molecular docking. European Review for Medical and Pharmacological Sciences, 24(8), 4529-4536. doi:10.26355/eurrev_202004_21036
Mahmud, S., Uddin, M.A.R., Zaman, M., Sujon, K.M., Rahman, M.E., Shehab, M.N., Islam, A., Alom, M.W., Amin, A., Akash, A.S., & Saleh, M.A. (2020). Molecular docking and dynamics study of natural compound for potential inhibition of main protease of SARS-CoV-2. Journal of Biomolecular Structure and Dynamics, Online ahead of print, 1-9. doi:10.1080/07391102.2020.1796808
Marwaha, A., Goel, R.K., & Mahajan, M.P. (2007). PASS-predicted design, synthesis and biological evaluation of cyclic nitrones as nootropics. Bioorganic and Medicinal Chemistry Letters, 17(18), 5251-5255. doi:10.1016/j.bmcl.2007.06.071
Morris, G.M., Huey, R., Lindstrom, W., Sanner, M.F., Belew, R.K., Goodsell, D.S., & Olson, A.J. (2009). AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of Computational Chemistry, 30(16), 2785-2791. doi:10.1002/jcc.21256
Mushtaq, S., Abbasi, B.H., Uzair, B., & Abbasi, R. (2018). Natural products as reservoirs of novel therapeutic agents. EXCLI Journal, 17, 420-451. doi:10.17179/excli2018-1174
Nakamura, K., Shimura, N., Otabe, Y., Hirai-Morita, A., Nakamura, Y., Ono, N., Altaf-Ul-Amin, M., & Kanaya, S. (2013). KNApSAcK-3D: a three-dimensional structure database of plant metabolites. Plant and Cell Physiology, 54(2), e4. doi:10.1093/pcp/pcs186
Pan, S.Y., Zhou, S.F., Gao, S.H., Yu, Z.L., Zhang, S.F., Tang, M.K., Sun, J.N., Ma, D.L., Han, Y.F., Fong, W.F., & Ko, K.M. (2013). New Perspectives on How to Discover Drugs from Herbal Medicines: CAM's Outstanding Contribution to Modern Therapeutics. Evidence-Based Complementary and Alternative Medicine, 2013, 627375. doi:10.1155/2013/627375
Prasanth, D.S.N.B.K., Murahari, M., Chandramohan, V., Panda, S.P., Atmakuri, L.R., & Guntupalli, C. (2020). In silico identification of potential inhibitors from Cinnamon against main protease and spike glycoprotein of SARS CoV-2. Journal of Biomolecular Structure and Dynamics, Online ahead of print, 1-15. doi:10.1080/07391102.2020.1779129
Pratama, M.R.F., Poerwono, H., & Siswodihardjo, S. (2020). Molecular Docking of Novel 5-O-benzoylpinostrobin Derivatives as SARS-CoV-2 Main Protease Inhibitors. Pharmaceutical Sciences, 26(Suppl 1), S63-S77. doi:10.34172/PS.2020.57
Sahidin, S., Salsabila, S., Wahyuni, W., Adryan, F., & Imran, I. (2019). Potensi Antibakteri Ekstrak Metanol dan Senyawa Aromatik dari Buah Wualae (Etlingera elatior). Jurnal Kimia Valensi, 5(1), 1-7. doi:10.15408/jkv.v5i1.8658
Sukardiman, S., Ervina, M., Pratama, M.R.F., Poerwono, H., & Siswodihardjo, S. (2020). The coronavirus disease 2019 main protease inhibitor from Andrographis paniculata (Burm.f) Ness. Journal of Advanced Pharmaceutical Technology and Research, 11(4), 157-162. doi:10.4103/japtr.JAPTR_84_20
Suwannarach, N., Kumla, J., Sujarit, K., Pattananandecha, T., Saenjum, C., & Lumyong, S. (2020). Natural Bioactive Compounds from Fungi as Potential Candidates for Protease Inhibitors and Immunomodulators to Apply for Coronaviruses. Molecules, 25(8), 1800. doi:10.3390/molecules25081800
Ulrich, S. & Nitsche, C. (2020). The SARS-CoV-2 main protease as drug target. Bioorganic and Medicinal Chemistry Letters, 30(17), 127377. doi:10.1016/j.bmcl.2020.127377
Wahyuni, W., Malik, F., Ningsih, A., Zubaydah, W.O.S., & Sahidin, S. (2018). Antimicrobial activities of ethanol extract of Wualae (Etlingera elatior (JACK) R.M. Smith). JPMS (Journal of Pharmaceutical and Medicinal Sciences). 3(1), 14-18.
World Health Organization. (2020). Coronavirus disease (COVID-19) Weekly Epidemiological Update and Weekly Operational Update: Weekly epidemiological update - 31 August 2020. https://www.who.int/publications/m/item/weekly-epidemiological-update---31-august-2020
Wu, C., Liu, Y., Yang, Y., Zhang, P., Zhong, W., Wang, Y., Wang, Q., Xu, Y., Li, M., Li, X., Zheng, M., Chen, L., & Li, H. (2020). Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharmaceutica Sinica B, 10(5), 766-788. doi:10.1016/j.apsb.2020.02.008
Zhu, L., George, S., Schmidt, M.F., Al-Gharabli, S.I., Rademann, J., & Hilgenfeld, R. (2011). Peptide aldehyde inhibitors challenge the substrate specificity of the SARS-coronavirus main protease. Antiviral Research, 92(2), 204-212. doi:10.1016/j.antiviral.2011.08.001
Authors
Authors continue to retain the copyright to the article if the article is published in the Borneo Journal of Pharmacy. They will also retain the publishing rights to the article without any restrictions.
Authors who publish with this journal agree to the following terms:
- Any article on the copyright is retained by the author(s).
- The author grants the journal, right of first publication with the work simultaneously licensed under a Creative Commons Attribution License that allows others to share work with an acknowledgment of the work authors and initial publications in this journal.
- Authors are able to enter into separate, additional contractual arrangements for the non-exclusive distribution of published articles of work (eg, post-institutional repository) or publish it in a book, with acknowledgment of its initial publication in this journal.
- Authors are permitted and encouraged to post their work online (e.g., in institutional repositories or on their websites) prior to and during the submission process, as can lead to productive exchanges, as well as earlier and greater citation of published work.
- The article and any associated published material are distributed under the Creative Commons Attribution-ShareAlike 4.0 International License.