In Silico Study to Assess Antibacterial Activity from Cladophora Sp. on Peptide Deformylase: Molecular Docking Approach
Abstract
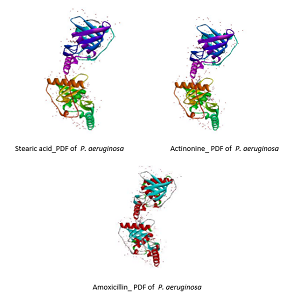
Increasing antibiotic-resistant pathogenic bacteria is a severe problem in the world. Therefore, there is a need to identify new drugs from natural products and also new drug targets. Cladophora sp. is a marine organism which is known to have bioactive compounds and a potential antibacterial. On the other hand, Peptide Deformylase (PDf) may prove to be a novel drug target since it is crucial for native peptide functioning in most pathogenic bacteria. This study screens for PDf inhibition activity of compounds from Cladophora sp. using molecular docking approach and screening the binding affinity of bioactive compounds against the peptide receptor PDf using Pyrex Autodock Vina software. Docking results were stored and visualized using Biovia Discovery Studio and PyMOL ligand. Ligands were obtained from previous literature in PubChem, and receptor peptide PDf from pathogenic bacteria: Pseudomonas aeruginosa (PDB ID:1N5N), Escherichia coli (PDB ID:1BSK), Enterococcus faecium (PDB ID:3G6N) and Staphylococcus aureus (PDB ID:1LQW), was obtained from the peptide data bank. The results of this screening show with ligand the highest binding affinity against PDf of P. aeruginosa, E. coli, E. faecium, and S. aureus is stearic acid (-5.9 kcal/mol), eicosapentaenoic acid (-6.6 kcal/mol), stearic acid (-5.8 kcal/mol), and stearic acid (-6.2 kcal/mol) respectively. The binding of natural compounds from Cladophora sp. with PDf models may provide a new drug with a different drug target for antibacterial potential.
Full text article
References
Apfel, C.M., Locher, H., Evers, S., Takacs, B., Hubschwerlen, C., Pirson, W., Page, M.G., Keck, W. 2001. Peptide deformylase as an antibacterial drug target: target validation and resistance development. Antimicrobial Agents and Chemotherapy. 45(4):1058-1064.
Centers for Disease Control and Prevention. 2013. Antibiotic Resistance Threats in the United States. Atlanta: Centers for Disease Control and Prevention.
Damayanti, D.S., Utomo, D.H., Kusuma, C. 2016. Revealing the potency of Annona muricata leaves extract as FOXO1 inhibitor for diabetes mellitus treatment through computational study. In Silico Pharmacology. 5(1):3.
Fieulaine, S., de Sousa, R.A., Maigre, L., Hamiche, K., Alimi, M., Bolla, J.M., Taleb, A., Denis, A., Pages, J.M., Artaud, I., Meinnel, T., giglione, C. 2016. A unique peptide deformylase platform to rationally design and challenge novel active compounds. Scientific Reports. 6:35429.
Gupta, P.K.P., Sahu, B. 2016. Identification of Natural Compound Inhibitors against Peptide Deformylase Using Virtual Screening and Molecular Docking Techniques. Bulletin of Environment, Pharmacology and Life Sciences. 4(9):70-80.
Munita, J.M., Arias, C.A. 2016. Mechanisms of Antibiotic Resistance. Microbiology Spectrum. 4(2): 10.1128/microbiolspec.VMBF-0016-2015.
Pratama, M.R.F., Pratomo, G.S. 2017. Pharmacophore Optimization of Berberine as HER2 Inhibitor. Jurnal Farmasi Indonesia. 14(2):109-117.
Pratama, M.R.F., Suhartono, E. 2018. Understanding the Interaction Between Glutathione and Acetaminophen: A Docking Study Approach. Dentino: Jurnal Kedokteran Gigi. 3(2):215-219.
Pratama, M.R.F., Suratno, S., Mulyani, E. 2018. Antibacterial Activity of Akar Kuning (Arcangelisia flava) Secondary Metabolites: Molecular Docking Approach. Asian Journal of Pharmaceutical and Clinical Research. 11(11):447-451.
Rampogu, S., Zeb, A., Baek, A., Park, C., Son, M., Lee, K.W. 2018. Discovery of Potential Plant-Derived Peptide Deformylase (PDF) Inhibitors for Multidrug-Resistant Bacteria Using Computational Studies. Journal of Clinical Medicine. 7(12):563.
Saadatmand, S., Khavarinejad, R., Nejadsattari, T., Soltani, S. 2011. Antioxidant and antibacterial activities of Cladophora glomerata (L.) Kütz. in Caspian Sea Coast, Iran. African Journal of Biotechnology. 10(39):7684-7689.
Snow-Snetzer, M., Sharifi-Rad, J., Setzer, W.N. 2016. The Search for Herbal Antibiotics: An In-Silico Investigation of Antibacterial Phytochemicals. Antibiotics. 5(3):E30.
Authors
Copyright (c) 2019 Yoni Rina Bintari, Rio Risandiansyah

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
Authors continue to retain the copyright to the article if the article is published in the Borneo Journal of Pharmacy. They will also retain the publishing rights to the article without any restrictions.
Authors who publish in this journal agree to the following terms:
- Any article on the copyright is retained by the author(s).
- The author grants the journal the right of first publication with the work simultaneously licensed under a Creative Commons Attribution License that allows others to share work with an acknowledgment of the work authors and initial publications in this journal.
- Authors can enter into separate, additional contractual arrangements for the non-exclusive distribution of published articles (e.g., post-institutional repository) or publish them in a book, with acknowledgment of their initial publication in this journal.
- Authors are permitted and encouraged to post their work online (e.g., in institutional repositories or on their websites) prior to and during the submission process. This can lead to productive exchanges and earlier and greater citations of published work.
- The article and any associated published material are distributed under the Creative Commons Attribution-ShareAlike 4.0 International License.